(First a quick update: Research Boost should now be running without any issues – we fixed the bug due to API outage of the latest Gemini model. Please let me know if you are still having issues for any reason. Now on to our regular scheduled post.)
Once upon a time, this is how my literature review looked like:
47 tabs open, 300+ PubMed hits, 12 “important-looking” PDFs on my desktop, and that sinking feeling in my gut that I’m still missing the one paper that matters.
But in the post-AI era (past 2 years), my workflow looks very different. While AI can help, it is important to know its drawbacks and keep the best from the past.
So now I use a hybrid approach:
- AI / Semantic Search: For the “quick and dirty” mapping of the territory.
- Traditional Keyword Search: For the deep dive and the gap-finding.
This post is about why I recommend this hybrid approach whether you are:
- trying to understand a topic,
- building the discussion section of a manuscript, or
- writing a full narrative literature review.
If you are doing a formal systematic review, this hybrid approach still helps, but with stricter rules (we’ll get there).
Keyword vs. AI-based / Semantic Search
Traditional lexical or keyword search, which relies on exact word matching, has been the go-to method for years. You type words into PubMed or Google Scholar and the engine looks for those words in the title, abstract, MeSH terms, etc.
As AI has matured, semantic search has emerged as a different way to think about “search” altogether. Instead of asking: “Which papers have these exact words?”, semantic search asks: “Which papers seem to be about this idea, even if the words differ?”
Both are useful. Both fail in predictable ways.
To design a better workflow, you need to understand how they think.
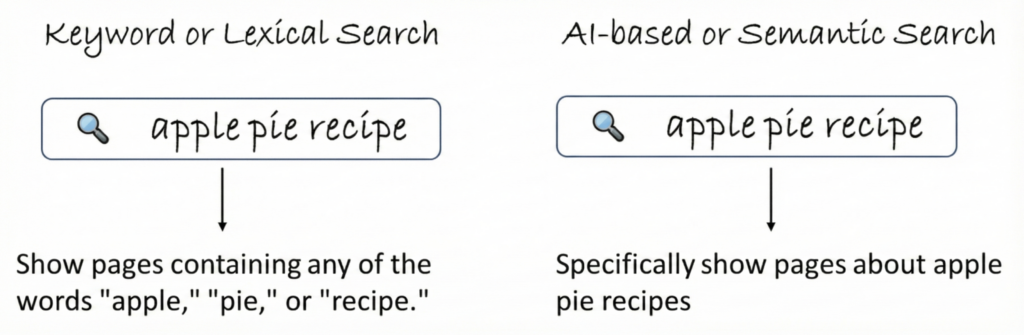
1. Keyword or Lexical Search: The Traditional Approach
Keyword/lexical search = literal matching.
You give PubMed a string like:
“psoriatic arthritis” AND “cardiovascular disease”
and it goes hunting for these exact tokens in selected fields (Title/Abstract, MeSH, etc.).
This has some clear advantages:
- High precision when you know the right words If your query is well-constructed, your hits are tightly focused.
- Reproducible Same string today and next year → same underlying search logic and results set (modulo newly indexed papers).
- Transparent You see the Boolean logic. You can debug it. You can show it in a PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) diagram.
But it also has obvious limitations:
- Vocabulary sensitivity If authors use “medication adherence,” “concordance,” “persistence,” or “compliance,” your search may or may not catch them depending on what you typed.
- Shallow understanding The engine doesn’t “know” that “heart attack” and “myocardial infarction” refer to the same clinical event unless you explicitly encode that with synonyms or MeSH.
In other words: keyword search is great when you already speak the language of the field. It punishes you when you don’t.
For classic keyword-based work, my defaults platforms are:
- PubMed as the baseline for most clinical and translational questions.
- Embase when I want more exhaustive coverage and meeting abstracts (especially useful for emerging topics).
- Scopus or Web of Science if I care about citation networks or broader interdisciplinary coverage.
- Google Scholar when I want a quick, messy but generous sweep of what’s out there.
Which ones you use will depend on how exhaustive you need to be, but PubMed + Embase + (optionally) Scopus already takes you a long way (Typically enough for even systematic reviews).
2. AI-Based or Semantic Search: The “Understands-Intent” Approach
AI-based or semantic search acts similar to a research assistant after a few months in your lab:
Even if you say it in slightly “wrong” language, they still understand what you mean.
Instead of matching raw words, semantic search:
- maps your question into a high-dimensional “meaning space” (vector embedding),
- compares that to embeddings of titles/abstracts/full texts,
- and retrieves content that is conceptually close to your query.
A few practical consequences for literature search:
- If you type: “intensive blood pressure lowering in people above 80”semantic search can still surface pivotal trial data such as that from HYVET and SPRINT subgroup analyses, even if they say “elderly,” “aged ≥80 years,” or “older adults” instead of “octogenarians.”
- If you type: “racial disparities in diabetes mortality”semantic search may retrieve studies on health inequities, socioeconomic determinants, or “outcome gaps”—even without the term “racial disparities.”
Under the hood, most modern semantic search systems rely on:
- Natural Language Processing (NLP) to parse and understand your question and the text of papers.
- Machine Learning to learn which results users tend to click or rate as useful, and adjust rankings over time.
- Sometimes knowledge graphs to link related entities (disease → drug class → outcome).
For us as researchers, the key point to understand is this:
Semantic search is optimized for intent and context, not just exact words.
It reduces the “you didn’t use the magic keyword, so we hid the paper from you” problem.
In terms of AI tools, my current semantic-search “stack” looks like this:
- Consensus – my top recommendation if you want a research-focused interface that summarizes and links directly to studies.
- SciSpace – very good for surfacing related papers and quickly understanding what a cluster of papers is about.
- Google Scholar’s new AI/semantic features – surprisingly useful now for quick, natural-language queries layered on top of the familiar Scholar index.
Tools like Litmaps and ResearchRabbit (visual/literature graph tools) are very popular and some people swear by them. Personally, I’ve found little added value for my own workflow beyond what a good keyword + semantic workflow gives me. If I really want an extensive search, I still reach for a carefully engineered keyword search rather than a fancy visual map—but I don’t discount that others may find those visual tools helpful for exploration or teaching.
How Keyword and Semantic Search Work Differently
| Aspect | Lexical / Keyword Search | AI / Semantic Search |
|---|---|---|
| Basis of Search | Exact strings, fields (Title/Abstract, MeSH) | Conceptual similarity in vector space |
| What it Optimizes For | Literal match, reproducibility | Contextual relevance, “are we talking about the same idea?” |
| Handling Synonyms & Jargon | Only if you manually include them (or via MeSH explosion) | Often automatic – understands that different phrases can point to the same concept |
| Relevance | Strong when query is carefully engineered | Strong even for messy, natural-language questions |
| Precision | Very high if query is expert-crafted | High at topic level, sometimes looser around edges |
| Recall (Coverage) | Risk of missing key papers if vocabulary is incomplete | Better coverage of “nearby” concepts and synonyms |
| Reproducibility | Stable. Same query → same core results | Variable. Rankings and included studies can shift between runs or tools. AI is non-deterministic by nature. |
| Hallucinations | None. You’re querying a curated index. | Risk exists if results are summarized by LLMs that can invent citations. This is getting better with state-of-the-art reasoning models. |
| User Experience | Demands familiarity with Boolean logic and domain terminology | Friendly for natural language, faster for “getting the lay of the land” |
Both types of search are just tools.
They answer different questions:
- Keyword search: “Given these exact terms, what does the database contain?”
- Semantic search: “Given this idea, what looks related even if it’s phrased differently?”
The Problem: The Precision vs. Recall Trap
Let me guess how your last lit review went.
Scenario A: The Keyword Trap
You started in PubMed. You typed your research question (e.g., “medication compliance in disease X”).
You got hits, but you missed a landmark study because the authors used:
- “medication adherence”
- or “concordance”
- or “persistence”
…and your keyword search was too literal to catch it.
- The Reality: Relying only on keywords means if you don’t know the exact password, you don’t get in.
Limitations of Traditional Keyword Search
Keyword search, which we have relied on for decades, isn’t perfect. It has its own set of limitations:
1. Over-reliance on hit counts
PubMed’s “534 results found” is not truth — it’s just what your exact string hits today. A small change in terms can swing results dramatically.
Takeaway: Hit counts are directional, not definitive.
2. Misses emerging or low-frequency terminology
Keyword tools privilege common vocabulary. New concepts, niche phenotypes, or early-phase studies may not appear at all unless you already know the exact term.
Takeaway: Keyword search overrepresents mature language and underrepresents emerging ideas.
3. No understanding of intent
“PsA prognosis” could refer to radiographic progression, functional decline, cardiovascular risk, or treatment response. Keyword search can’t distinguish.
Takeaway: It matches your words, not your meaning.
4. High-volume phrases create noise
Terms like “quality of life,” “outcomes,” or “compliance” retrieve highly heterogeneous literature, inflating workload without improving relevance.
Takeaway: Keywords can look academically important but bring mostly noise.
5. No sense of importance
Keyword matching treats a landmark RCT and a one-line case series with equal weight if the terms appear.
Takeaway: Keyword search reports presence, not priority.
Scenario B: The AI Trap
So, you swung to the other extreme. You opened an AI tool and asked:
“What is the prevalence of X in patients with Y?”
Boom. A fluent summary. It feels amazing for 30 seconds.
Then the doubt creeps in. Did it miss the key trial because it’s a “black box”?
- The Reality: AI is non-deterministic. A study on the AI tool Elicit found that the same query gave 246, 169, and 172 results in three different runs. While not all literature review needs to be systematic, one would hope that we don’t miss pivotal studies. And that is a real possibility if that pivotal study is hidden inside a paywall. If it is invisible to the AI, it might not exist.
- The other problem that I see commonly is that if there are multiple meeting abstracts related to your question, it might provide more weight to those (open-access) compared to peer-reviewed papers which might be under a paywall. The meeting abstracts are typically much longer (400 to 500 words) compared to the 250 to 300-word limit for a manuscript abstract, which may be the only thing the AI can see for that paper.
Common Failure Modes of AI-Based or Semantic Search
Semantic search is powerful, but it has predictable blind spots:
1. Messy or incomplete data → messy results
Semantic models only “understand” what they can see. If abstracts are sparse, metadata is incorrect, or journals are poorly indexed, the embeddings become noisy. Conference abstracts (long, open-access) often look more “relevant” than paywalled full papers simply because they contain more text.
Takeaway: Poor or incomplete data → unreliable semantic relevance.
2. Wrong model for the job
Many embedding models are trained on general web text, not clinical language. So terms like “MI,” “RA,” or “progression” can be misunderstood or mapped to irrelevant concepts.
Takeaway: If the model isn’t biomedical-tuned, it may misinterpret your query without you ever seeing the error.
3. Ranking ≠ clinical importance
Semantic similarity is not the same as clinical relevance. AI might rank lengthy abstracts, older guidelines, or tangentially related papers above the key RCTs if ranking logic is simplistic or opaque.
Takeaway: “Conceptually close” does not always mean “clinically important.”
Putting It Together: A Fair Comparison
At this point, here’s the mental model I actually use and teach:
| Feature | Traditional Keyword Search (PubMed/MeSH) | AI / Semantic Search (Vector/LLM) |
|---|---|---|
| How it “thinks” | Literal strings + Boolean logic | Conceptual similarity, intent, context |
| Strengths | Reproducible, auditable, essential for SLRs | Great for exploration, discovering vocabulary, mapping a field |
| Main Failure Modes | Misses synonyms, overfocus on popular terms, can’t see intent or importance | Sensitive to data quality, model choice, ranking defaults, paywall blindness |
| Hallucination Risk | None | Non-zero if you rely on generated citations/summaries |
| Best Role | Final evidence retrieval, confirmatory search | Early “scout” and vocabulary teacher |
So both ends of the spectrum have risks.
Once you think about it this way, the hybrid workflow becomes almost obvious.
My Pipeline for AI-Based Search
Conceptually, most semantic search tools go through a similar pipeline:
- Understand your question. The system uses natural language processing (NLP) to parse your query: “intensive BP lowering in octogenarians” → recognizes “intensive BP lowering,” elderly population, outcome of interest.
- Embed your question + documents into the same “meaning space.” Each abstract or paper is turned into a vector. Your question is also turned into a vector. The system looks for vectors that are close.
- Rank and refine. Machine learning models reorder results based on similarity scores plus whatever ranking signals the tool designers chose (click behavior, journal reputation, recency, etc.).
You don’t see any of this. What you see is a ranked list that, ideally, “feels right” conceptually.
But you still have to remember:
- you don’t fully control what corpus it can see,
- you don’t fully control how it ranks,
- and you can’t easily reproduce the same run months later for PRISMA.
That’s why I never stop at semantic search.
I use it as a scout, not as the final arbiter of truth.
The Hybrid Workflow
I realized I wanted the speed and vocabulary help of AI with the rigor and reproducibility of PubMed.
Also, there is research that shows AI search may find articles that are missed by classic keyword search. In the study on the AI tool Elicit, it found three additional papers that had been missed by the classic screening method.
So the hybrid workflow brings in the best of both worlds.
Here is the workflow I actually use for narrative reviews, grant backgrounds, and idea validation.
STEP 1: The AI “Scout” (AI-based search)
Goal: Identify 5–10 “signal” papers and learn the language of the field.
I start with a broad, natural-language question in a semantic search tool.
My Input: “Does intensive blood pressure lowering benefit octogenarians?”
What the semantic engine does well:
- It doesn’t care whether the title says “octogenarians,” “elderly,” or “aged ≥80.”
- It tends to surface conceptually related, higher-signal trials (SPRINT, HYVET, key meta-analyses).
Why this is useful:
- Semantic tools overweight abstracts, which is exactly what we want at this point: a quick mapping of the terrain.
- I’m not trying to exhaustively capture every subgroup analysis yet. I just want to know: What are the main pillars of this field?
Think of this step as “let the AI teach me the vocabulary and main trials.”
In other words, it significantly accelerates the discovery process.
STEP 2: The “Distill” Phase
Goal: Extract the actual language, entities, and structures the field uses.
Here, I am not “reading deeply.” I’m mining.
From those 5–10 signal papers, I pay attention to:
- Population phrasing
aged 80 and over,very elderly,older adults, not just “octogenarians.”
- Intervention phrasing
intensive blood pressure lowering,tight control of hypertension,SBP < 120 mmHg.
- Named entities
- Trial acronyms (
HYVET,SPRINT), - Key outcomes (
stroke,all-cause mortality,cardiovascular events), - MeSH terms appearing in indexing.
- Trial acronyms (
Now my goal shifts:
From “What is going on in this field?”
to “How do people in this field name what is going on?”
Once I have their vocabulary, I can return to a tool that likes clear, rigid syntax: PubMed (plus Embase/Scopus/others depending on how broad I need to go).
STEP 3: The “Drill Down” (Targeted Keyword Search)
Goal: Build a rigorous, reproducible evidence base.
Now I translate those terms into a structured search strategy.
Example:
(“hypertension”[Title/Abstract] OR “high blood pressure”[Title/Abstract])
AND (“tight control”[Title/Abstract] OR “intensive therapy”[Title/Abstract])
AND (“aged, 80 and over”[MeSH] OR “very elderly”[Title/Abstract] OR “HYVET”[Title/Abstract])
What I usually see:
- Precision improves. The key trials from Step 1 are all here.
- Recall improves in a controlled way. I now capture additional subgroup analyses, follow-up papers, and related work that the AI did not surface—but that match my carefully tuned keywords and MeSH.
- Reproducibility is restored. If a co-author asks “How did you get this set of 57 papers?” I can show the exact string.
In practice, I often end up with ~50–60 high-quality, directly relevant papers in about 10–12 minutes using this Scout → Distill → Drill Down loop.
Why We Automated This into Research Boost
For the past 2 years, I had been teaching how to do this manually.
- We copied and pasted DOIs into spreadsheets.
- We manually scanned abstracts to spot MeSH terms and trial acronyms.
- We built Boolean queries by hand and de-duplicated references in Excel or a reference manager.
At some point it hit me:
This is a system. Systems can be automated.
So, inside Research Boost, we codified this exact manual approach:
- Semantic Scout You ask a natural-language question. We run a semantic search to find candidate “signal” papers.
- Strategy Extraction The tool scans those papers and pulls out recurring keywords, MeSH terms, trial acronyms, and other entities.
- The Drill Down Research Boost auto-builds PubMed-style queries from these terms and runs them against real databases.
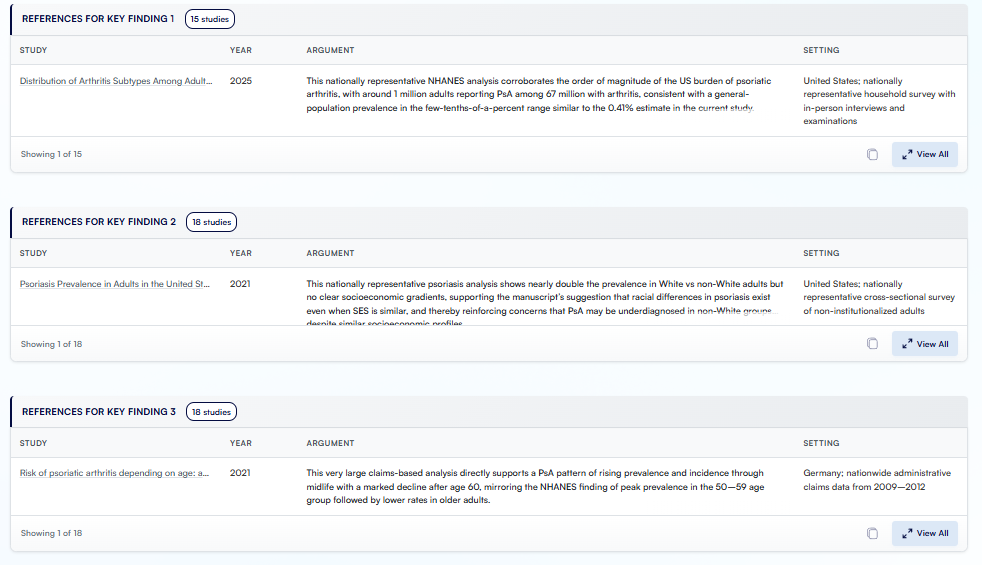
Figure. A snapshot of the literature review that Research Boost did for the discussion section.
You still stay in control:
- You can verify each of the papers by clicking on the links.
- Review the argument and relevant snippet for each of the papers.
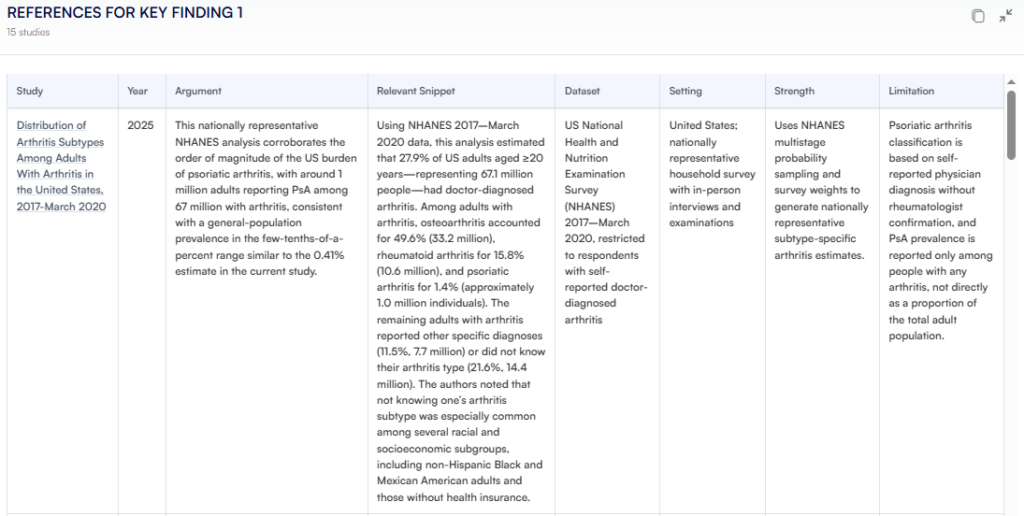
Figure. Details of each study that Research Boost provides.
Because Research Boost ultimately retrieves text from PubMed/journal sites, rather than relying on the model’s “memory,” we have not encountered hallucinated references across hundreds of tests so far. The databases act as the fact-checkers; the AI helps with framing, distilling, and connecting.
The Exception: What About Systematic Reviews?
If you are doing a formal Systematic Review or Meta-analysis with a PRISMA diagram, you have stricter constraints.
For those projects:
- AI-based retrieval as the primary source of studies is a NO-GO.
- Non-determinism and opaque ranking mean you cannot guarantee that the same query will give the same pool of papers in the future.
However, AI is still incredibly useful before you run the “official” search:
- You can use semantic search to learn vocabulary, synonyms, MeSH terms, typical inclusion/exclusion criteria.
- You then build a transparent, reproducible search strategy and run it solely in PubMed, Embase, Cochrane, etc.
You don’t use the AI to find the final set of studies.
You use the AI to do an initial “feasibility” search and to narrow or solidify your research question.
It can also help you write the code (the Boolean query) that you then execute in traditional databases. You can copy and paste the below prompt:
The “Systematic Review Strategist” Prompt
If you want a reproducible PubMed string and don’t want to manually brainstorm 50 synonyms, here is the exact prompt I use.
You can plug this into your favorite LLM (ChatGPT, Gemini, Claude, etc.):
You are an academic literature search expert, who collaborates and helps researchers and physician-scientists in an academic institution.
Take a deep breath and let’s do this step-by-step.
Step 1: Produce a list of 50 relevant terms.
Follow my instructions precisely to develop a highly effective Boolean query for a medical systematic review literature search. Do not explain or elaborate. Only respond with exactly what I request.
First, given the following research question or topic, identify 50 terms or phrases that are relevant. The terms you identify should be used to retrieve more relevant studies, so be careful that the terms you choose are not too broad. Do not include duplicates.
Statement: [INSERT YOUR RESEARCH QUESTION HERE]
Step 2: Classify terms using PICO
For each item, label it as one of: terms relating to health conditions (A), terms relating to a treatment (B), terms relating to types of study design (C). If it doesn’t fit, mark as (N/A). Each item must be categorised into (A), (B), (C), or (N/A).
Step 3: Create a Boolean Query in PubMed syntax.
Using the categorised list, create a Boolean query that can be submitted to PubMed which groups items from each category, e.g.:
((itemA1[Title/Abstract] OR itemA2[Title/Abstract]) AND (itemB1[Title/Abstract] OR itemB2[Title/Abstract]) AND (itemC1[Title/Abstract] OR itemC2[Title/Abstract]))
Step 4: Refine search strategy and add MeSH.
Refine the query to retrieve as many relevant documents as possible while controlling volume. Add relevant MeSH terms where appropriate, e.g., MeSHTerm[MeSH]. Retain the general structure, with each main clause corresponding to a PICO element. Ensure the final query is valid PubMed syntax.
Use that to design your search. Then run it in PubMed yourself.
Closing Thought
We’re in a transition period.
You don’t have to choose between:
- being “old-school” and living only in PubMed, or
- being “AI-native” and trusting whatever a chatbot tells you.
Use the hybrid method.
- For narrative reviews, grant backgrounds, and scoping work: Let an AI-augmented workflow (or a tool like Research Boost) automate the Scout → Distill → Drill Down loop.
- For systematic reviews and any context where you must show reproducibility: Use AI to help design the search strategy, but let PubMed (and friends) do the actual retrieval.
That’s how you keep your work fast, modern, and still grounded in reality.
For your next review, don’t obsess over being “pure” about methods.
Design a workflow that respects your time and still respects the science.
PROMPT OF THE WEEK
The Grant Ripple Effect Analyzer
Prompt:
I’m considering **[clearly describe the grant decision you’re evaluating—e.g., adding/removing a Specific Aim, changing the primary hypothesis, shifting scope, choosing a different mechanism, modifying budget/personnel, resubmitting versus creating a new grant, etc.]**.
Help me think beyond the obvious, the way an experienced study section reviewer would.
1. **Map the first-order effects**
Identify the immediate, direct consequences for:
* Significance, Innovation, and Approach
* Feasibility within the budget and timeline
* Fit with the chosen funding mechanism and FOA/RFA
* My role and the team’s credibility (PI/multi-PI, environment, collaborators)
2. **Analyze the second-order effects**
Describe indirect, follow-on consequences for:
* Perceived coherence of the Aims page and overall narrative
* Risk of over-ambitious scope versus “too incremental” science
* Alignment with review criteria (e.g., clarity of hypotheses, rigor, reproducibility, human subjects protections)
* Downstream demands on data, methods, and required preliminary work
3. **Explore the third-order effects**
Think through longer-term ripple impacts that applicants often overlook, such as:
* How this decision shapes my future funding trajectory and niche
* Synergy or conflict with existing grants and pending applications
* Implications for trainees, collaborations, and institutional politics
* How reusable the work/products will be for future grants and papers
4. **Highlight risks, unintended consequences, and hidden dependencies**
Surface:
* Methodologic or feasibility risks that reviewers are likely to flag
* Dependencies on resources, datasets, or personnel that could fail
* Any misalignment between aims, methods, budget, and timeline
* Ethical, regulatory, or compliance issues that could derail the application
5. **Suggest alternate choices or mitigations**
Propose:
* Alternative framings, aim structures, or scope adjustments
* Ways to de-risk the proposal while preserving innovation
* Concrete mitigations I can bake into the Approach, timeline, or budget
* Brief example reviewer comments I might receive under different choices
Think like a senior grant reviewer who cares about fundability, coherence, and long-term programmatic impact, not just getting this one application submitted.
Now ask me about my grant and the change I want to make.
P.S. My good friend, Razia Aliani, is running an accredited 12-hour workshop on “High-Efficiency Systematic Reviews (with Ethical AI Integration)” for researchers who want to conduct Cochrane-standard SRs with ethical, transparent AI support. Razia is a senior systematic review consultant, Cochrane-trained, whose thesis focused entirely on AI tools in SRs long before ChatGPT. This program is built on real projects for academia, NGOs, government agencies, and pharma.
Participants learn end-to-end SR methodology, ethical AI workflows, modern screening strategies, live demos of the best tools, plus eBooks, templates, scripts, and practical resources to work faster without compromising rigor.
You can get an exclusive 30% off with code PARAS30 (valid until Dec 7):
🔗https://systematicreview.razia-aliani.com
(I do not get any % of this → I have loved her content on the same and would highly recommend.)
